Research
Research in our group concerns the enzymology of histidine biosynthesis and tRNA methylation. We employ a breadth of strategies and techniques (e.g. gene cloning and expression, site-directed mutagenesis, protein purification, steady-state and pre-steady-state kinetics, isothermal titration calorimetry, differential scanning fluorimetry, mass spectrometry, NMR, isotopic labelling of substrates and enzymes, kinetic and binding isotope effects, density-functional theory calculations, protein crystallography, inhibitor design and synthesis) to elucidate the mechanisms of enzymatic reactions. We harness that information to design specific enzyme inhibitors of promising novel targets for antibiotic and anticancer development.
ATP phosphoribosyltransferase and phosphoribosyl-ATP pyrophosphohydrolase/phosphoribosyl-AMP cyclohydrolase
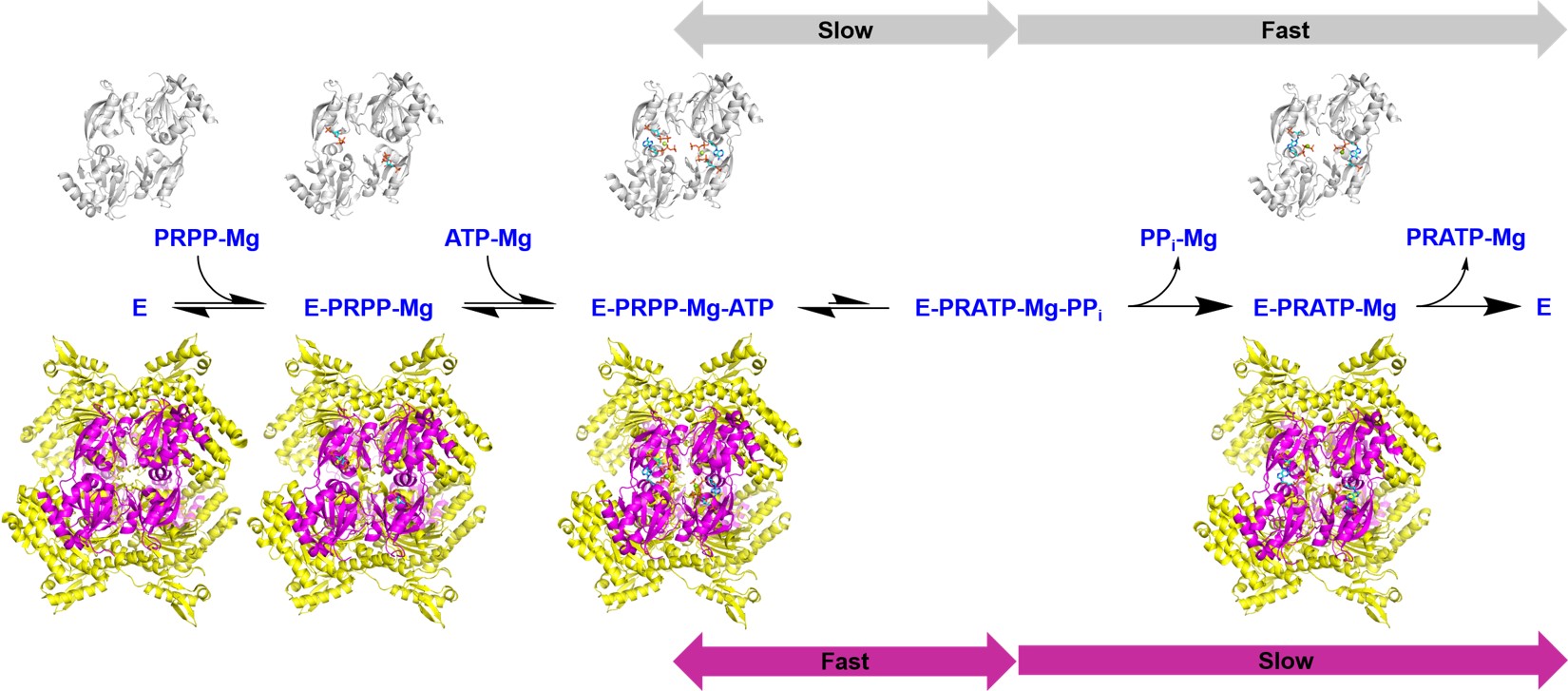 The biosynthesis of histidine involves some unusual enzymatic reactions. Efforts are focused on the first, second and third steps of the pathway, the hetero-octameric and allosteric ATP phosphoribosyltransferase (ATPPRT) and the bifunctional phosphoribosyl-ATP pyrophosphohydrolase/phosphoribosyl-AMP cyclohydrolase (HisIE). The long-term goal is to understand the principles that underpin enzyme catalysis and allostery in these systems to enable inhibitor design towards novel antibiotics against Acinetobacter baumannii.
The biosynthesis of histidine involves some unusual enzymatic reactions. Efforts are focused on the first, second and third steps of the pathway, the hetero-octameric and allosteric ATP phosphoribosyltransferase (ATPPRT) and the bifunctional phosphoribosyl-ATP pyrophosphohydrolase/phosphoribosyl-AMP cyclohydrolase (HisIE). The long-term goal is to understand the principles that underpin enzyme catalysis and allostery in these systems to enable inhibitor design towards novel antibiotics against Acinetobacter baumannii.
m1A22-tRNA methyltransferase
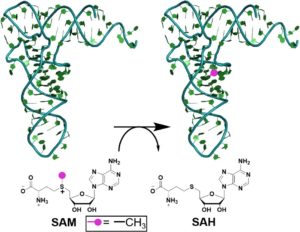 The enzyme m1A22-tRNA methyltransferase (TrmK) is essential for Staphylococcus aureus survival but absent in humans. We are elucidating the mechanism and substrate specificity of this enzyme, and developing covanlent inhibitors of TrmK towards novel antibiotics against MRSA. We are also investigating human tRNA methyltransferases involved in resistance to current cancer treatments.
The enzyme m1A22-tRNA methyltransferase (TrmK) is essential for Staphylococcus aureus survival but absent in humans. We are elucidating the mechanism and substrate specificity of this enzyme, and developing covanlent inhibitors of TrmK towards novel antibiotics against MRSA. We are also investigating human tRNA methyltransferases involved in resistance to current cancer treatments.